I recently had the pleasure of attending the 10th Gateway Computing Environments (GCE) Workshop. The GCE program was created about 10 years ago as part of NSF TeraGrid project. Nancy Wilkins-Diehr of the San Diego Supercomputing Center has been the leader of GCE from day one and she continues to do an amazing job with the project and organizing the workshops. GCE has become a large and very productive collaboration in the U.S. and there is also now an International branch with meetings in Europe and Australia.
If you are unfamiliar with the science gateway concept a one paragraph tutorial is in order. A science gateway is a web portal or app that is tailored to research needs of a specific scientific community. The gateway can allow its users access to advanced applications and data collections. For example, the Cyberinfrastructure for Phylogenetic Research (CIPRES) gateway (http://www.philo.org) explores the evolutionary relationships among various biological species. It allows its users to run dozens of computationally intensive phylogenetic tree inference tools (e.g., GARLI, RAxML, MrBayes) on NSF’s XSEDE supercomputers. Another great example is NanoHub that was built at Purdue University to provide education and research tools for nanotechnology. In the genomics research world GenomeSpace is a gateway for a large collection of tools and it represents a broad community of developers and users.
I was around for the early days of GCE and I also attended the International event in Dublin in 2014. I have been amazed by the progress this community has made. Ten years ago the science gateway program in TeraGrid was a small project that was seen by the big computing centers as “mostly harmless”[1]. However it is now a big deal. NanoHub has evolved into Hubzero.org and it supports many additional domains including cancer research, pharmaceutical manufacturing, earthquake mitigation, STEM education. They have had 1.8 million users in 2014. CIPRES is now one of XSEDE’s largest users consuming 18.7M compute core hours in 2014. Of course there are many big individual users doing great science at scale on the NSF supercomputer centers, but CIPRES and some other gateways represent about one half of all XSEDE users. The bottom line is that the nation’s investment in supercomputing is paying off for a very wide spectrum of scientists.
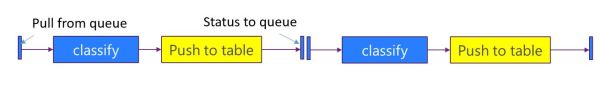
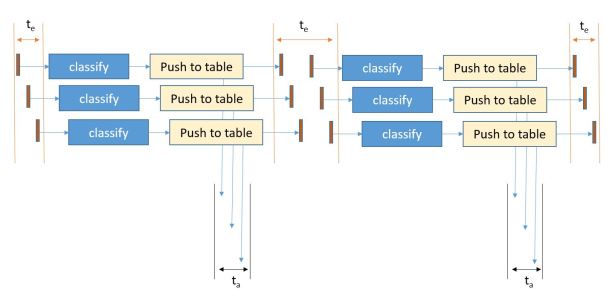
The workshop had some excellent talks. There was a fantastic Keynote by Alexandra Swanson from Oxford and the Zooniverse project. Zooniverse is a rather unusual gateway. Rather than being supercomputer powered it is “people powered”. It started with the Galaxy Zoo project but now supports just about any research project that needs large teams of volunteers to help sift through vast piles of (mostly) image data and help with classification and discovery. It is a bit like the Amazon mechanical Turk but much more science focused. Ali’s talk (slides here) really stressed the challenge of building and designing the portal technology need to allow a user to build a productive instance of a universe.
The slides for all the presentations are on the program page for the workshop. They were all good but two of them were very interesting to me. Pankaj Saha from Binghamton University gave a great talk about how they are now integrating Docker, Marathon and Mesos with the Apache Airavita project. Airavita is a framework for orchestrating the computational jobs and workflows of a science gateway onto supercomputers, local clusters and commercial clouds. It is high component based and a natural fit for Docker, Marathon and Mesos. The talk was only a snapshot of work in progress and I look forward to seeing how well they are able to exploit these tools to make Airavita extremely easy to deploy and manage.
Shava Smallen of the SDSC gave a fascinating talk about how machine learning can be used to predict failures and manage resources for the CIPRES gateway. My experience with one science gateway many years ago was that various parts were always crashing. (At the time, I preferred to blame the users and not our software.) This SDSC project is really interesting. I am convinced that this is a great application for ML. Distributed system software like Zookeeper, Mesoshpere and Marathon along with extremely reliable messaging services like RabbitMQ have made huge strides in our ability to design robust systems. But Supercomputing platforms are designed to push the edge in terms of performance and are not very friendly to long running systems like science gateways. Doing more of this sort of “smart” monitoring and failure prediction is very important.
Sustainable Open Source Scientific Software
The other Keynote at the event was by Dan Katz from Chicago and NSF. While his talk was not about science gateways he did raise an issue that is of great concern to the gateway community. Specifically how do we maintain software when the grant funding that paid for its construction runs dry? This question has been around for as long as scientists have been writing code.
I think we have to ask the following question. What makes open source software sustainable over long periods of time? In other words, who pays the salary of the programmers tasked with maintaining and improving it? I believe there are several answers to these questions. Fundamentally when a piece of software becomes a critical component of the infrastructure of an entity or entities with deep pockets those entities will make sure that the software is sustained. There are a number of big tech companies that depend upon various bits of open source and they pay their people to make sure it continues to work. I know that Microsoft is an active contributor to several open source platforms. Companies like IBM, Facebook, Twitter, Amazon, and Google are all supporting open source projects. Often these same companies will take something that was developed internally and open source it because they see value in attracting other contributors. Good examples are Yahoo and Hadoop and Google and Kubernetes. Other sustaining entities are organizations like CERN, the National Institute of Health, the Department of Energy and NASA. But in these cases the software being sustained is critical to their specific missions. The NSF has a greater challenge here. Its budget is already loaded with supporting more critical infrastructure than it can afford if it is to maintain its mission to support cutting edge science across a vast spectrum of disciplines.
The other way open source software originating in universities is sustained is to spin-off a new start-up company. But to make this happen you need angel or venture funding. That funding will not appear if there is no perceived value or business model. Granted that in today’s start-up frenzy value and business model are rather fuzzy things. Often these companies will find markets or they will be bought up by larger companies that depend upon its product. UC Berkeley has been extremely impressive in this regard. DataBricks.com has spun out of the successful Spark data analysis system and Mesosphere.com is a spin-off form the Mesos project. Both of these are very exciting and have a bright future.
The third way open source survives is the labor of love model. Linux was maintained by dedicated cadre of professionals, many of whom contribute in the off-hours after their day jobs. The Python community is another remarkable example and the best example of sustaining open source scientific software. Companies like Enthought and Continuum Analytics have emerged from the community to support Python for technical computing. Enthought supports SciPy.org, the great resource for scientific Python tools. Continuum provides the peerless free Anaconda python package. And the Numfocus foundation has been created by the community to channel financial sponsorship to many of the most significant science projects of interest to the Python user community.
Looking at the success of the Python, Hadoop, Mesos and Spark one can see critical common threads that are missing in many research software projects. First there must be a user community that is more than casual. It must be passionate. Second it must be of value to more than these passionate scientists. It must help solve business problems. And this is the challenge facing the majority of science software which is often too narrowly focused on a specific science discipline. I have often seen groups of researchers that have the critical mass to make a difference, but they are not sufficiently collaborating. How many different scientific workflow systems do we need? All too often the need for academic or professional promotion gets in the way of collaboration and the result is duplication.
I believe the Science Gateway community is doing the right thing. I see more consolidation of good ideas into common systems and that is great. Perhaps they should create a Numfocus-like foundation? And they need to solve some business problems. 🙂
[1] A description borrowed from the Hitchhikers Guide to the Galaxy. The full description for the planet Earth was “mostly harmless”. This was the revised extended entry. The earlier entry for the planet Earth was “harmless”.